Jimena Golcher-Benavides - Does population genetic divergence predict biodiversity patterns?
Jimena Golcher-Benavides
Biodiversity Graduate Student Research Enhancement Grant Awardee
https://jimenagolcher.wordpress.com/
Program in Ecology (PiE) and Department of Botany, University of Wyoming
Graduate Advisor: Dr. Catherine E. Wagner
Introduction
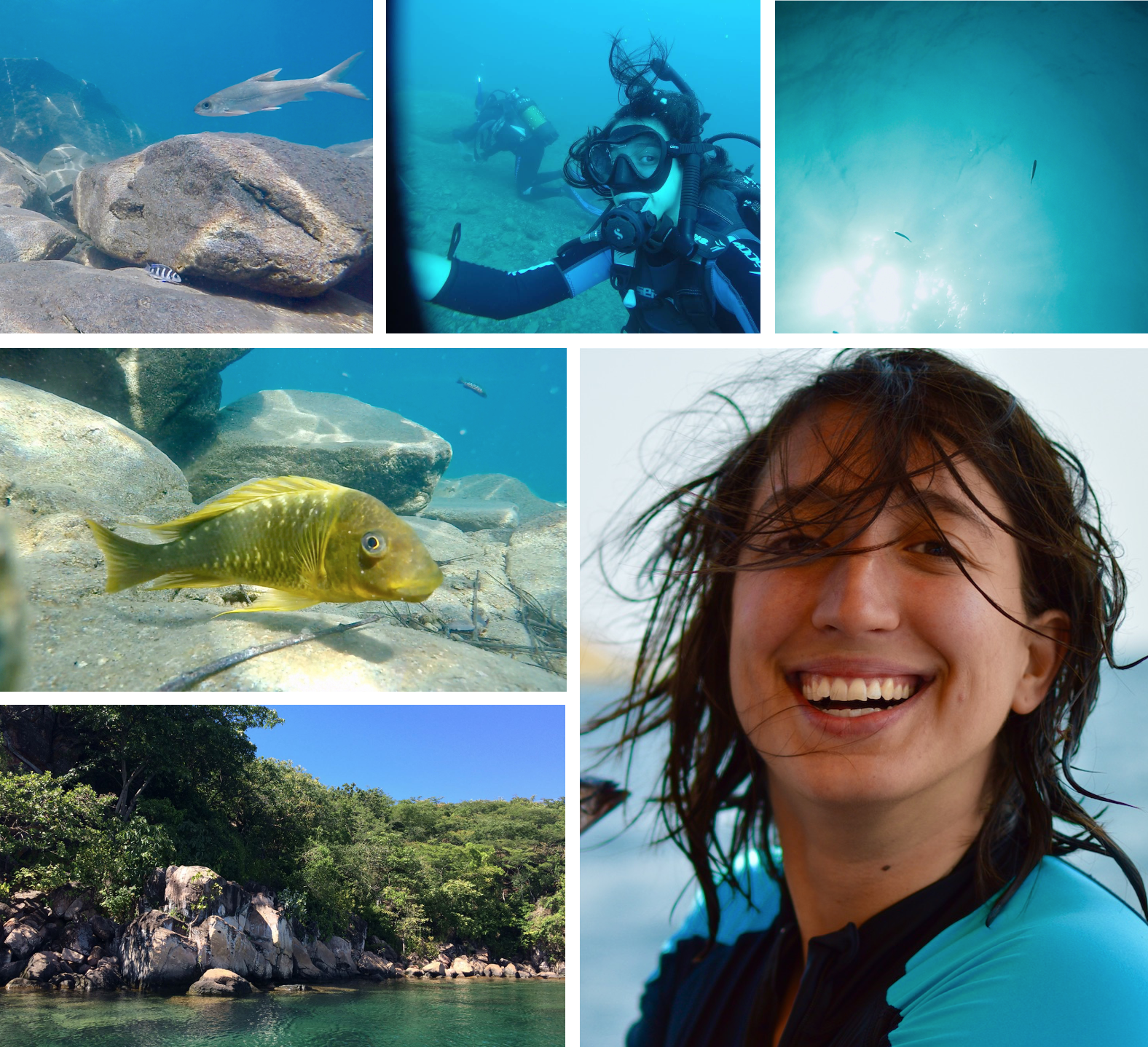

Objective of the project
Although many studies have sought to test for links between traits and diversification rates e.g., relatively few studies have explored the relationship between populationlevel processes and macroevolutionary patterns e.g. In the effort to link population-level processes to speciation, and speciation patterns to diversification, a key goal is better understanding the links between organismal traits and population-level demography (e.g. dispersal rates; population size). Patterns in spatial decay in genetic similarity (known as isolation-by-distance, or IBD) occur because of dispersal limitations leading to greater likelihood of individuals mating with individuals that are close in geographically rather than distant. Dispersal limitation may result from: (1) extrinsic environmental factors affecting animal movement (e.g. dispersal barriers), or extrinsic drivers governing population growth dynamics (e.g. resource availability), and (2) intrinsic organismal traits limiting the probability for dispersal (e.g. traits influencing movement). Studying patterns of spatial decay in genetic similarity in both species-rich and species-poor lineages, and the correlates of these spatial patterns, will help us to identify the drivers underlying hotspots and coldspots in biodiversity.
Share This Page
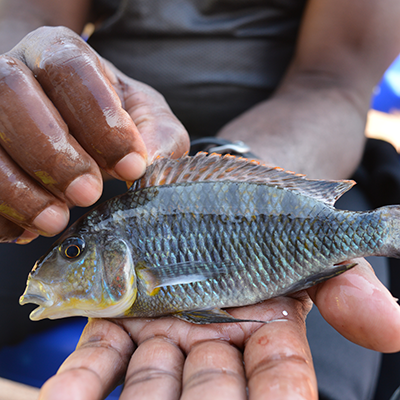
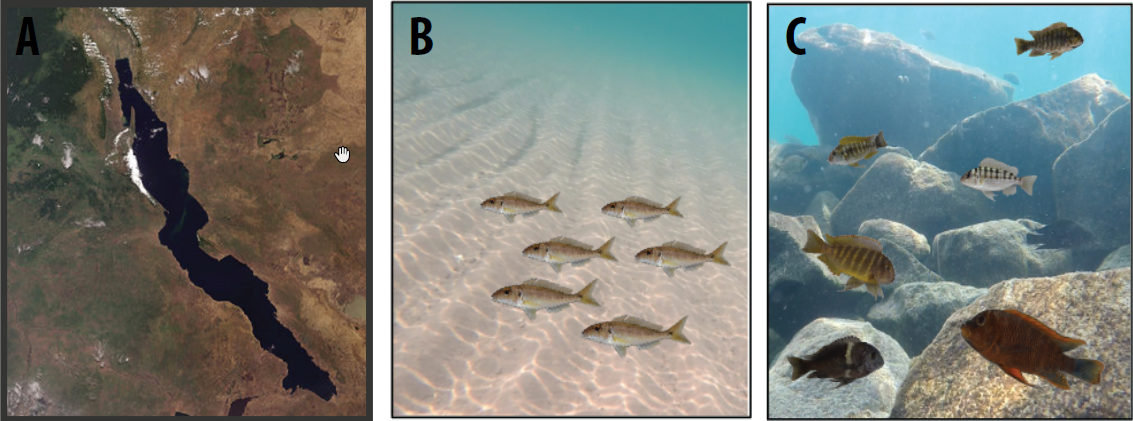


Fig. 1: Satellite view of Lake Tanganyika, East Africa (A), cichlid diversity in the sand and in the rocky littoral zone (B-C).
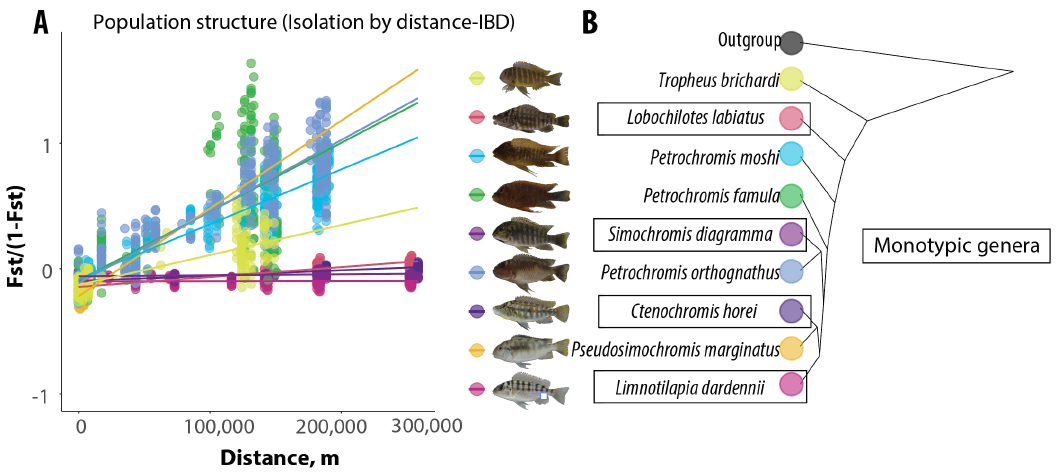
Fig. 2: Preliminary data on ß slopes calculated for 4 monotypic genera and 5 non-monotypic taxa from species-rich genera (A). A cladogram depicting relationships among taxa is shown in B. These preliminary data indicate that species from monotypic genera show less population structure than nonmonotypic genera.
Research Highlights





